 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
Compute the buried surface area of two given partners. More...
#include <Buried_surface_area.hpp>
Public Types | |
| typedef T_Alpha_complex_of_molecular_model< ParticleTraits > | Alpha_complex |
| Derived class from CGAL alpha-complex for filling it with particles. | |
Compute the buried surface area of two given partners.
Compute the buried surface area of two given partners
| ParticleTraits | Model of the concept ParticleTraits. |
| FT | Number type used for representing the buried surface area (default is intervals of double). |
| typedef T_Alpha_complex_of_molecular_model<ParticleTraits> Alpha_complex |
Derived class from CGAL alpha-complex for filling it with particles.
It uses ParticleTraits for determining the geometrical kernel used for the representation and computation of the 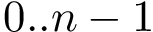
| ParticleTraits | Traits class defining geometric properties of a particle. |
| TAlphaComplexVertexBase | Base class for representing a vertex in the 3D |
| TAlphaComplexCellBase | Base class for representing a cell in the 3D |
| TAlphaComplexBase | Base template representation of the |