 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
Compute the Betti numbers of a 3D Cell Complex from its alpha-complex. More...
#include <Betti_numbers_2.hpp>
Public Types | |
| typedef AlphaComplex3::FT | FT |
| Number type used for representing coordinates and weights in the | |
| typedef boost::tuple< unsigned, unsigned, unsigned > | result_type |
| Boost tuple (Betti_0, Betti_1, Betti_2) | |
Functors | |
| result_type | operator() (const AlphaComplex3 &A) const |
| Returns the Betti numbers of the input | |
| template<class InputIterator> | |
| result_type | operator() (InputIterator begin, InputIterator end, const FT &alpha=0) const |
| Computes the Betti numbers of the union of the family of input 3D balls represented as weighted points. | |
Compute the Betti numbers of a 3D Cell Complex from its alpha-complex.
| AlphaComplex3 | A Model of the 3D |
Number type used for representing coordinates and weights in the 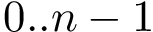
| typedef boost::tuple<unsigned, unsigned, unsigned> result_type |
Boost tuple (Betti_0, Betti_1, Betti_2)
|
inline |
Returns the Betti numbers of the input 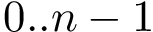
|
inline |
Computes the Betti numbers of the union of the family of input 3D balls represented as weighted points.