Nearest_neighbors_graph_builder
Authors: F. Cazals and T. Dreyfus and C. Roth
Introduction
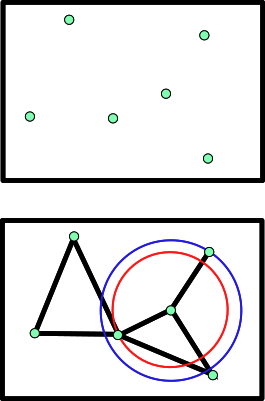
This package provides operations to build a nearest neighbors graph (NN graph for short), basically a Boost Graph from the BGL library, with weights on the edges representing distances.
The NN graph may be defined by linking all vertices to a subset of its neighbors, k closest or in a given range r. All proimity queries rely on an underlying spatial search engine, as explained in the package Spatial_search .
Implementation
Builders
Consider a Boost Graph G from the BGL library such that each vertex is associated to a point. To be precise, it is assumed that the bundle property of the vertex is a point. The package provides a concept of NearestNeighborsGraphBuilder for building such a graph from a range of points [begin,end) and a spatial search engine (search_engine).
In the following, we assume that one is given a search engine either returning the neighbors of a sample within a distance range, or returning the k nearest neighbors of a sample. Three operations are defined:
- 1) inserting a range of point [begin, end) into a graph G – one vertex is created for each point;
NearestNeighborsGraphBuilder(begin, end, G);
- 2) connecting a vertex u in G to its neighbors, using a search engine :
NearestNeighborsGraphBuilder(u, search_engine, G);
- 3) repeating the last operation for every vertices of G :
NearestNeighborsGraphBuilder(search_engine, G);
The package provides two models of builders :
- SBL::GT::Nearest_neighbors_graph_builder_with_edge_weight , a builder assuming that the type of NNG handles weights over the edges, so that distances between vertices bounding an edge can be stored. This is particularly useful when the distance function is heavy to compute, so that it is only computed during the build operation of the NNG.
Module
This package provides the module SBL::Modules::T_Nearest_neighbors_graph_builder_module< ModuleTraits > to build a NNG from an input set of points. The template parameter ModuleTraits must define three types :
- Nearest_neighbors_graph : the type representing the NNG, derived from the Boost Graph library;
- Nearest_neighbors_distance : the distance functor between two points (it has to define itself the types Point representing a point, and FT representing the coordinates type;
- Nearest_neighbors_query< Distance > : a template class for the spatial search engine, parametrized by a distance functor between vertices of the NNG;
The main options of the module T_Nearest_neighbors_graph_builder_module are:
–num-neighbors int: Target number of neighbors for each vertex.
–distance-range int: Distance range specification.
–nng-sym bool: Symmetric NNG ie connects two vertices iff they are mutual nearest neighbors.
–nng-connected int: Attempt connecting the NNG by iteratively increasing the number of neighbors; halt if NNG is connected, or if the prescribed num of neighbors is reached.
–nng-file int: Skip calculations by loading the NNG from a file.
Examples
Simple builder
This example shows how to build the NNG of 2D points built on 100x100 grid, setting four neighbors per point.
#include <SBL/GT/Nearest_neighbors_graph_builder.hpp>
#include <SBL/GT/ANN_metric_space_proximity_tree.hpp>
#include <SBL/GT/ANN_meta.hpp>
#include <iostream>
class Point_type
{
public:
double x;
double y;
Point_type(void) : x(0), y(0){}
Point_type(const Point_type& p) : x(p.x), y(p.y){}
Point_type(double i, double j) : x(i), y(j){}
friend std::ostream& operator<<(std::ostream& out, const Point_type& p)
{
out << "[" << p.x << " " << p.y << "]";
return out;
}
};
typedef boost::adjacency_list<boost::vecS, boost::vecS, boost::undirectedS, Point_type> Graph;
typedef boost::graph_traits<Graph>::vertex_descriptor Vertex;
class Distance
{
public:
typedef double FT;
typedef Vertex Point;
Graph* m_G;
Distance(void):m_G(NULL){}
Distance(Graph& G):m_G(&G){}
FT operator()(Vertex p, Vertex q)const
{
return sqrt(((*this->m_G)[p].x - (*this->m_G)[q].x)*((*this->m_G)[p].x - (*this->m_G)[q].x) + ((*this->m_G)[p].y - (*this->m_G)[q].y)*((*this->m_G)[p].y - (*this->m_G)[q].y));
}
};
int main(){
std::vector<Point_type> points;
for(double i = 0; i < 100; i++)
for(double j = 0; j < 100; j++)
points.push_back(Point_type(i, j));
Graph G;
NNG_builder builder;
builder.compute_sym = true;
builder(points.begin(), points.end(), G);
Distance dist(G);
ANN query(dist);
query.set_query_type(ANN::ANN_BY_K);
query.set_number_of_neighbors(4);
query.insert(boost::vertices(G).first, boost::vertices(G).second);
builder(query, G);
std::cout << "G has " << boost::num_vertices(G) << " vertices and "
<< boost::num_edges(G) << " edges." << std::endl;
return 0;
}
Builder for nearest neighbors graphs.
Definition Nearest_neighbors_graph_builder.hpp:64
Nearest neighbor engine designed for metric spaces.
Definition ANN_metric_space_proximity_tree.hpp:101
Builder with edge weights
This example performs the same task as previous but with a NNG able to store the distances between pairs of points linked bya nedge in he NNG.
#include <SBL/GT/Nearest_neighbors_graph_builder.hpp>
#include <SBL/GT/ANN_metric_space_proximity_tree.hpp>
#include <SBL/GT/ANN_meta.hpp>
#include <iostream>
class Point_type
{
public:
double x;
double y;
Point_type(void) : x(0), y(0){}
Point_type(const Point_type& p) : x(p.x), y(p.y){}
Point_type(double i, double j) : x(i), y(j){}
friend std::ostream& operator<<(std::ostream& out, const Point_type& p)
{
out << "[" << p.x << " " << p.y << "]";
return out;
}
};
typedef boost::property<boost::edge_weight_t, double> Edge_property;
typedef boost::adjacency_list<boost::vecS, boost::vecS, boost::undirectedS, Point_type, Edge_property> Graph;
typedef boost::graph_traits<Graph>::vertex_descriptor Vertex;
class Distance
{
public:
typedef double FT;
typedef Vertex Point;
Graph* m_G;
Distance(void):m_G(NULL){}
Distance(Graph& G):m_G(&G){}
FT operator()(Vertex p, Vertex q)const
{
return sqrt(((*this->m_G)[p].x - (*this->m_G)[q].x)*((*this->m_G)[p].x - (*this->m_G)[q].x) + ((*this->m_G)[p].y - (*this->m_G)[q].y)*((*this->m_G)[p].y - (*this->m_G)[q].y));
}
};
int main(){
std::vector<Point_type> points;
for(double i = 0; i < 100; i++)
for(double j = 0; j < 100; j++)
points.push_back(Point_type(i, j));
Graph G;
NNG_builder builder;
builder(points.begin(), points.end(), G);
Distance dist(G);
ANN query(dist);
query.set_query_type(ANN::ANN_BY_K);
query.set_number_of_neighbors(4);
query.insert(boost::vertices(G).first, boost::vertices(G).second);
builder(query, G);
std::cout << "G has " << boost::num_vertices(G) << " vertices and "
<< boost::num_edges(G) << " edges." << std::endl;
return 0;
}
Builder for nearest neighbors graphs.
Definition Nearest_neighbors_graph_builder.hpp:122


