 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
Authors: S. Marillet and P. Boudinot and F. Cazals
This package provides tools to (i) select binding affinity prediction models from atomic coordinates, typically crystal structures of the partners and/or of the complex, and to (ii) exploit such models to perform affinity predictions.
At the selection stage, given a set of predefined variables coding structural and biochemical properties of the partners and of the complex, the approach consists of building sparse regression models which are subsequently evaluated and ranked using repeated cross-validation [124] .
At the exploitation stage, the variables selected are used to build a predictive model on a training dataset, and predict the affinity on a test dataset.
More precisely, the approach runs through 3 steps:
Consider two species ![]() and
and ![]() forming a complex
forming a complex ![]() . The dissociation is defined by
. The dissociation is defined by ![]() , namely the ratio between the concentration of individual partners and bound partners.
, namely the ratio between the concentration of individual partners and bound partners.
The strength of this association is quantified by the dissociation free energy ![]() , which, in the
, which, in the ![]() standard state, satisfies with
standard state, satisfies with ![]() Boltzmann's constant and
Boltzmann's constant and ![]() the temperature:
the temperature:
This equation shows that ![]() has two components respectively coding the enthalpic (
has two components respectively coding the enthalpic ( ![]() ), and entropic (
), and entropic ( ![]() ) changes upon binding. As stated above, we wish to estimate these quantities from atomic coordinates, typically crystal structures of the partners and/or the complex.
) changes upon binding. As stated above, we wish to estimate these quantities from atomic coordinates, typically crystal structures of the partners and/or the complex.
Parameters
The parameters used in this package are based on the following key geometric constructions:
Using the previous notions, we define four categories of atoms of a complex:
Biophysical rationale.
Before defining our parameters, we raise several simple observations underlying their design:
Using the quantities defined in section General pre-requisites, we define the following variables:
Interface terms.
Packing terms.
Consider the volume variation of ![]() , see Eq. (eq-atom-volume-variation). As recalled above, packing properties are important in several respects. Adding up volume variations yields the following four sum of volumes differences (VD) parameters:
, see Eq. (eq-atom-volume-variation). As recalled above, packing properties are important in several respects. Adding up volume variations yields the following four sum of volumes differences (VD) parameters:
Solvent interactions and electrostatics.
The interaction between a protein molecule and water molecules is complex. In particular, the exposition to the solvent of non-polar groups hinders the ability of water molecules to engage into hydrogen bonding, yielding an entropic loss for such water molecules. We define the following terms:
Parameters used to estimate binding affinities.
There are three groups of parameters (see the next equations for their definition):
The acronyms read as follows (see text for details):
For variables describing a change between the bound and unbound forms of the partners e.g. ![]() or
or ![]() , it is necessary to match the atoms of the bound partners to those of the unbound partners. For this, the amino-acid sequences of the partners are extracted from the PDB files and aligned. Then, the atoms of every pair of corresponding residues are matched using their name.
, it is necessary to match the atoms of the bound partners to those of the unbound partners. For this, the amino-acid sequences of the partners are extracted from the PDB files and aligned. Then, the atoms of every pair of corresponding residues are matched using their name.
Since missing residues and missing atoms are common in crystal structures, the proportion of matched atoms is computed as to investigate potential wrong matchings, and discard entries for which the alignment is not good enough.
In this section we explain how to compute the variables presented in section Associated variables .
This step consists of two sub-steps:
The input of the pre-processing step consists in:
An XML file complying with the following format:
Step 1.
The first step of the workflow is performed by the script ![]() which is called as follows:
which is called as follows:
sbl-bap-step-1-run-applications.py -i data/example_data.xml -s data/eisenberg_solvation_parameters.txt -p data/structures -n 1 -d results
| File Name | Description |
| Plain text annotations file | Eisenberg Solvation Parameters |
| XML file | PDB ids and chains of complexes and unbound partners |
| PDB file | Example PDB file in the input directory of option -p |
Step 2.
The second step of the workflow is performed by the script ![]() which is called as follows:
which is called as follows:
sbl-bap-step-2-compile-molecular-data.py -f data/example_data.xml -d results -o results/affinity_dataset.xml
Step 1.
Practically, this first step consists in running ![]() (through SBL::PDB_complex_to_unbound_partners_matchings::PDB_complex_to_unbound_partners_matcher ),
(through SBL::PDB_complex_to_unbound_partners_matchings::PDB_complex_to_unbound_partners_matcher ), ![]() and
and ![]() on various entries of the dataset. For each entry, a directory named after the following scheme is created in the current working directory: <PDB_ID>_<chains_A>_<chains_B>
on various entries of the dataset. For each entry, a directory named after the following scheme is created in the current working directory: <PDB_ID>_<chains_A>_<chains_B>
Each directory is structured as follows:
1A2K_AB_C |-- 1OUN_A_A_atom_matchings.txt |-- 1OUN_A_A_residue_matchings.txt |-- 1OUN_AB | |-- sbl-vorlume_eisenberg_solvation_parameters__log.txt | |-- sbl-vorlume_eisenberg_solvation_parameters__surface_volumes.xml |-- 1OUN_B_A_atom_matchings.txt |-- 1OUN_B_A_residue_matchings.txt |-- 1QG4_A | |-- sbl-vorlume_eisenberg_solvation_parameters__log.txt | |-- sbl-vorlume_eisenberg_solvation_parameters__surface_volumes.xml |-- 1QG4_C_A_atom_matchings.txt |-- 1QG4_C_A_residue_matchings.txt |-- sbl-vorlume_eisenberg_solvation_parameters__log.txt |-- sbl-vorlume_eisenberg_solvation_parameters__surface_volumes.xml |-- sbl-vorshell-bp-ABW-atomic__AB_ball_shelling_forest.dot |-- sbl-vorshell-bp-ABW-atomic__AB_ball_shelling_forest.xml |-- sbl-vorshell-bp-ABW-atomic__AB_packing.xml |-- sbl-vorshell-bp-ABW-atomic__BA_ball_shelling_forest.dot |-- sbl-vorshell-bp-ABW-atomic__BA_ball_shelling_forest.xml |-- sbl-vorshell-bp-ABW-atomic__BA_packing.xml |-- sbl-vorshell-bp-ABW-atomic__interface_AB.pdb |-- sbl-vorshell-bp-ABW-atomic__interface_BA.pdb |-- sbl-vorshell-bp-ABW-atomic__log.txt |-- sbl-vorshell-bp-ABW-atomic__surface_volumes.xml
This example is for entry 1A2K with partners AB and C. All *_matchings.txt files contain the matchings between atoms or residues of the bound and unbound structures. Directories 1OUN_AB and 1QG4_A are for the unbound partners. All sbl-* files contain the results of the SBL executables. If verbose output is toggled (-v flag), various information will be output on the standard and error output.
Step 2.
| Preview | File Name | Description |
| XML archive | Compiled results of runs of step 1 |
In this section, we explain how to select a binding affinity model maximizing a performance criterion, using the script ![]() . Two points worth noticing are:
. Two points worth noticing are:
In the sequel, we explain how to predict ![]() of complexes from a dataset
of complexes from a dataset ![]() , using the methodology summarized on Fig fig-bap-workflow . Estimation is performed on a per complex basis, from which performances at the whole dataset level are derived.
, using the methodology summarized on Fig fig-bap-workflow . Estimation is performed on a per complex basis, from which performances at the whole dataset level are derived.
Our predictions rely on three related concepts defined precisely hereafter (see also Fig fig-bap-workflow):
Templates.
Denote ![]() the pool of variables. Let a template be a set of variables, i.e. a subset of
the pool of variables. Let a template be a set of variables, i.e. a subset of ![]() . To define parsimonious templates from the set
. To define parsimonious templates from the set ![]() , we generate subsets of
, we generate subsets of ![]() involving up to at most
involving up to at most ![]() variables. This defines a pool of templates
variables. This defines a pool of templates ![]() .
.
Cross-validation.
In the following a model is associated to both a template ![]() and a dataset
and a dataset ![]() . More precisely, a model refers to a regression model, i.e. the variables of the template plus the associated coefficients.
. More precisely, a model refers to a regression model, i.e. the variables of the template plus the associated coefficients.
Practically, models are defined during ![]() -fold cross-validation, and a number of
-fold cross-validation, and a number of ![]() of repetitions (see Fig fig-bap-workflow). Consider one repetition, which thus consists of splitting at random
of repetitions (see Fig fig-bap-workflow). Consider one repetition, which thus consists of splitting at random ![]() into
into ![]() subsets called folds. For one fold, a regression model associated with
subsets called folds. For one fold, a regression model associated with ![]() is trained on (k-1)/k of the dataset
is trained on (k-1)/k of the dataset ![]() , and predictions are run on the remaining 1/k of complexes. Processing the
, and predictions are run on the remaining 1/k of complexes. Processing the ![]() folds yields one repetition of the cross validation procedure, resulting in one prediction
folds yields one repetition of the cross validation procedure, resulting in one prediction ![]() for the
for the ![]() of each complex. The set of all predictions in one repeat, say the
of each complex. The set of all predictions in one repeat, say the ![]() th one, is denoted
th one, is denoted
Note again that these predictions stem from ![]() regression models associated with
regression models associated with ![]() , namely one per fold.
, namely one per fold.
Statistics per template.
Considering one cross-validation repetition ![]() , we define the correlation
, we define the correlation ![]() as the correlation between the experimental values
as the correlation between the experimental values ![]() and the predictions
and the predictions ![]() . An overall assessment of the template
. An overall assessment of the template ![]() using the
using the ![]() repetitions is obtained by the following median of correlations
repetitions is obtained by the following median of correlations
For a complex ![]() , we define the binding affinity prediction
, we define the binding affinity prediction ![]() as the median across repetitions i.e.
as the median across repetitions i.e.
The median prediction error is defined by
and the median absolute prediction error by:
Using this latter value, we define the prediction ratio ![]() as the percentage of cases such that the dissociation free energy is off by a specified amount
as the percentage of cases such that the dissociation free energy is off by a specified amount ![]() :
:
In particular, setting ![]() to 1.4, 2.8 and 4.2 kcal/mol in the previous equation yields cases whose
to 1.4, 2.8 and 4.2 kcal/mol in the previous equation yields cases whose ![]() is approximated within one, two and three orders of magnitude respectively.
is approximated within one, two and three orders of magnitude respectively.
Finally, a permutation test yields a p-value for each predictive model [144] .
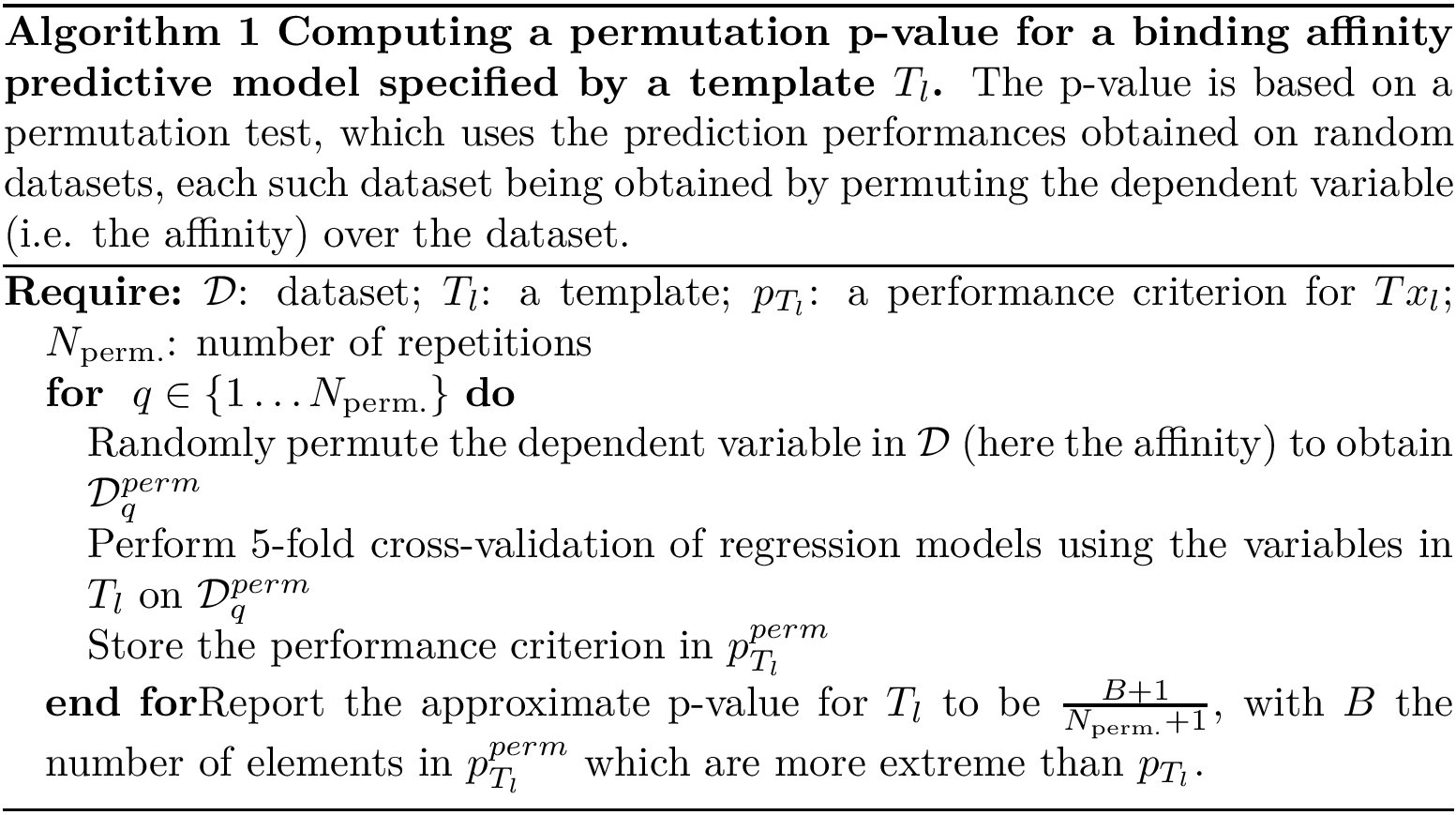
In a nutshell, the rationale consists of generating randomized datasets by shuffling their ![]() values. Then, one computes the performance criterion for each such dataset, from which the p-value is inferred (see Algorithm~alg-p-val).
values. Then, one computes the performance criterion for each such dataset, from which the p-value is inferred (see Algorithm~alg-p-val).
Model selection.
Define the best predictive model as the one maximizing performance criterion. We wish to single out the best predictive models, i.e. those that cannot be statistically distinguished from the best predictive model. More precisely, consider a pool of predictive models ![]() , out of which we wish to identify a subset of predictive models whose distribution cannot be distinguished from that of the best predictive model. To this end, let H0 be the null hypothesis stating that two predictive models yield identical performance distributions. We decompose the predictive models as
, out of which we wish to identify a subset of predictive models whose distribution cannot be distinguished from that of the best predictive model. To this end, let H0 be the null hypothesis stating that two predictive models yield identical performance distributions. We decompose the predictive models as ![]() such that :
such that :
The predictive models in ![]() are called the specific models for the dataset
are called the specific models for the dataset ![]() . The corresponding procedure is based on the Kruskal-Wallis test, see [124] .
. The corresponding procedure is based on the Kruskal-Wallis test, see [124] .
The p-value threshold for this test is set to ![]() .
.
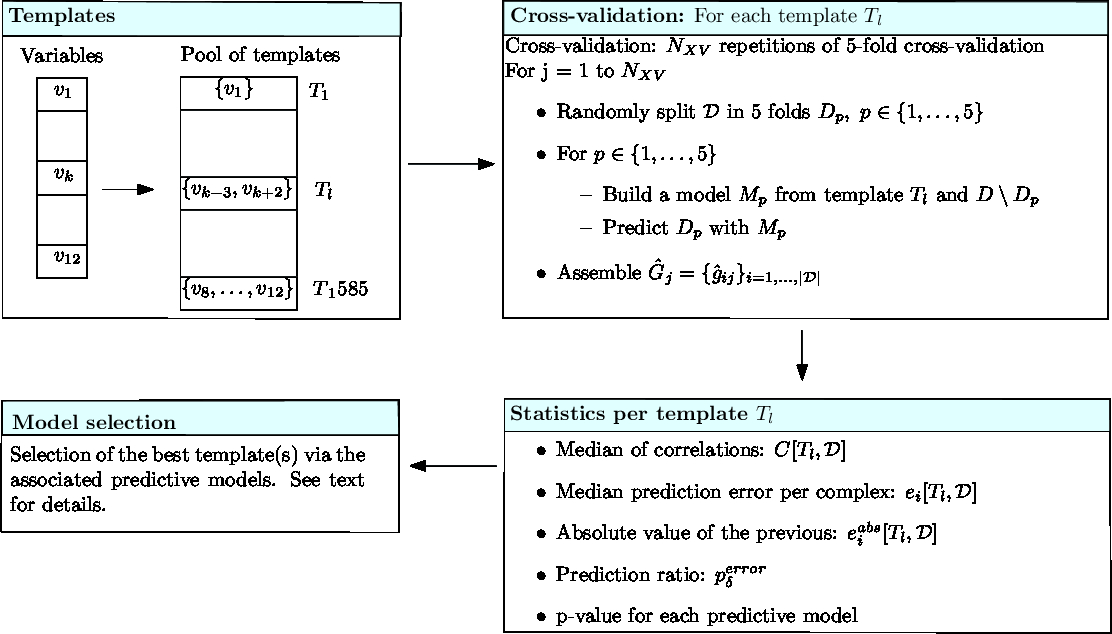 |
Running binding affinity predictions for a dataset (Cross-validation) Each template undergoes a number (Statistics) Various statistics are computed to assess the performances yielded by the predictive model associated to each template. (Model selection) Predictive models are compared, and the best ones selected. |
Model generation and selection is performed by the script ![]() which is executed as follows :
which is executed as follows :
# Quick example: maximum 2-variables models, 10 repetitions and p-value permutations, # flexible complexes only (irmsd > 1.5 A), run on a single process sbl-bap-step-3-models-analysis.py -f results/affinity_dataset.xml -m 2 -n 1 -r 10 -p 10 -l 1.5 -o flex_quick
# Heavier example: maximum 4-variables models, 100 repetitions and # 1000 p-value permutations, all complexes, run on three # processes. Performs k-nearest neighbors regression with 10 neighbors sbl-bap-step-3-models-analysis.py -f results/affinity_dataset.xml -m 4 -n 3 -r 100 -p 1000 -k 10 -o all_heavier
The XML file specified by option -f should comply with the following format:
The txt file specified by option -s should comply with the following format:
| File Name | Description |
| XML archive file | Compiled data generated from |
This step outputs a minimum of seven files:
Files 1 and 2 are text files with the following structure:
In the first one, the index and the ranks are computed on the cross-validated median absolute error of the model. In the second they are computed on the median cross-validation correlation coefficient.
Files 3 and 4 below contain a matrix of prediction per cross-validation repetition for the best model. Here "best" refers to the model selected by the model selection procedure described above. If multiple models are selected, each one has an associated file. These files are structured as follows:
File 5 and 6: PDF files containing scatter plots of the prediction made by the best predictive models (from files 1 and 2 respectively) for each complex against the experimental affinity value.
| Preview | File Name | Description |
| Plain text file | File 1 | |
| Plain text file | File 2 | |
| Plain text file | File 3 | |
| Plain text file | File 4 | |
| PDF file | Scatter plot of the prediction versus experimental affinity for the best model(s) | |
| PDF file | Scatter plot of the prediction versus experimental affinity for the best model(s) | |
| PDF file | Scatter plot of the prediction error versus interface RMSD |
Once a set of variable has been selected during the previous step, it can be used to estimate a model. Recall that a model is a function returning an estimate for binding affinity of a complex. For this, one needs a training set and a regressor, i.e. an object able to use the training set to predict the affinity of a new dataset (for instance, linear least-squares fitting, ![]() nearest neighbors, regression trees, ...). An example of the procedure to train such a regressor on the whole Structure Affinity Benchmark and predict its own entries (that is to perform predictions without cross-validation) is given in a demo file (sbl-binding-affinity-model-exploitation-example.py).
nearest neighbors, regression trees, ...). An example of the procedure to train such a regressor on the whole Structure Affinity Benchmark and predict its own entries (that is to perform predictions without cross-validation) is given in a demo file (sbl-binding-affinity-model-exploitation-example.py).
We now describe two algorithms used by this package. The first simply computes an approximate (upper bound on the) permutation p-value for a given statistic [144] . The pseudo-code is given on Algorithm alg-p-val .
The second algorithm is used to divide a pool of models into two groups, namely ![]() , as discussed in the section Using the programs to select affinity prediction models . The corresponding procedure is based on the Kruskal-Wallis test, and can be found in [124] .
, as discussed in the section Using the programs to select affinity prediction models . The corresponding procedure is based on the Kruskal-Wallis test, and can be found in [124] .

We describe successively the 2 steps of section Using the programs to pre-process structural data .
Step 1.
The script ![]() runs applications from the SBL on the bound and unbound partners.
runs applications from the SBL on the bound and unbound partners.
The matching between atoms (section Matchings for atoms) is done using the python module SBL::PDB_complex_to_unbound_partners_matchings which is a wrapper around the executable ![]() . This executable performs the matching using the class SBL::CSB::T_Alignment_engine_sequences_seqan, from the package Alignment_engines, which itself uses the Seqan library to perform the sequence alignment, and subsequently match residue and atoms based on the alignment found.
. This executable performs the matching using the class SBL::CSB::T_Alignment_engine_sequences_seqan, from the package Alignment_engines, which itself uses the Seqan library to perform the sequence alignment, and subsequently match residue and atoms based on the alignment found.
In order to compute shelling orders and packing properties, the executables ![]() and
and ![]() are also run on the complex, and
are also run on the complex, and ![]() is run on the unbound partners.
is run on the unbound partners.
Step 2.
The script ![]() compiles the previous results and computes the variables listed in section Associated variables for each entry of the dataset.
compiles the previous results and computes the variables listed in section Associated variables for each entry of the dataset.
Data structures to store the properties of entities such as atoms, residues, chains and files are grouped in the python module SBL::Protein_complex_analysis_molecular_data_structures . To fill these data structures it also provides a class used to traverse the directory structure and parse the files resulting from step 1.
The machinery used to build, evaluate and rank various models using the previously computed variables can be found in the python module SBL::Combinations_of_variables_model_evaluation::Combinations_of_variables_model_evaluation .
Once a set of variables has been selected, model fitting on a training set and prediction of a new dataset can be done using any software or language. An example script is given using python and scikit-learn in file ![]()
The seqan library [73] (BSD, Seqan) is necessary when computing the variables which encode the variations between bound and unbound partners.