 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
Packages | |
| Module_base | |
 | Framework for building generic blocks of algorithms and generic workflows for applications. Reference Manual – User Manual . Authors: F. Cazals and T. Dreyfus |
Classes | |
| class | T_Archive_file_loader< InputArchive, SerializedData > |
Loader for one or more boost archive files More... | |
| class | T_Multiple_archives_xml_archive_file_loader< SecondaryArchive, SecondaryData, SerializedData > |
Loader for one or more multiple archives xml archive files More... | |
| class | T_Primitive_labels_loader< PartnerLabelsTraits, MediatorLabelsTraits, ExtraLabelsTraits > |
| Loader for systems' labels specification. Loader for systems' labels specification. More... | |
| class | T_PDB_file_loader< ESBTLMolecularSystem, PDBLineFormat > |
| Loader for one or more PDB files, even listed in a file. Loader for one or more PDB files, even listed in a file. More... | |
| class | T_Points_d_file_loader< PointD > |
| Loader for one or more txt files listing dD points Loader for one or more txt files listing dD points. More... | |
| class | T_Spheres_3_file_loader< Sphere3, Point3 > |
Loader for one or more txt files listing 3D spheres More... | |
| class | T_Alignment_sequences_module< ModuleTraits, AlignmentEngineSequences > |
| Module which computes a pairwise alignment of two sequences Module which computes a pairwise alignment of two sequences. More... | |
| class | T_Alignment_structures_module< ModuleTraits, AlignmentEngineStructures > |
| Module which computes a pairwise alignment of two structures Module which computes a pairwise alignment of two structures. More... | |
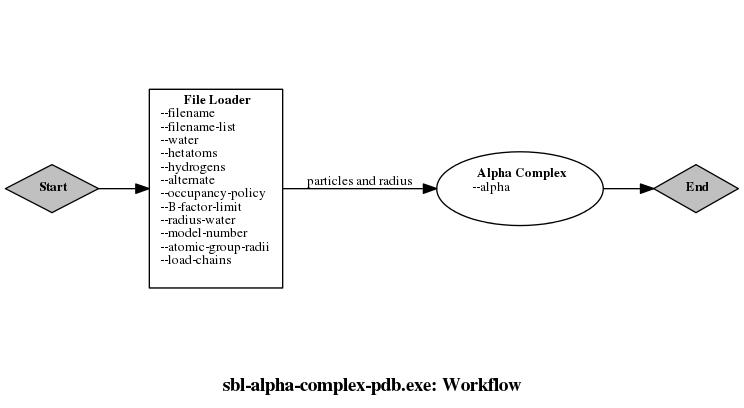
| class | T_Alpha_complex_of_molecular_model_module< ModuleTraits > |
| Module building the alpha-complex of an input set of particles. Module building the alpha-complex of an input set of particles. More... | |
| class | T_Buried_surface_area_with_labels_module< ModuleTraits > |
| Module building the buried surface areas from the alpha-complex of a molecular model. Module building the buried surface areas from the alpha-complex of a molecular model. More... | |
| class | T_Buried_surface_area_without_label_module< ModuleTraits > |
| Module building the buried surface area of a partner from the alpha-complex of a molecular model. Module building the buried surface area of a partner from the alpha-complex of a molecular model. More... | |
| class | T_Cluster_engine_module< ModuleTraits > |
| Module which computes clusters from a cloud of points Module which computes clusters from a cloud of points. More... | |
| class | T_Minimal_oritend_spanning_forest_module |
| Module building the minimal spanning forest of a weighted bipartite graph Module building the minimal spanning forest of a weighted bipartite graph. More... | |
| class | T_Molecular_interfaces_module< ModuleTraits > |
| Module building the bicolor, mediated and tricolor interfaces of a molecular structure. Module building the bicolor, mediated and tricolor interfaces of a molecular structure. More... | |
| class | T_Molecular_structure_classifier_module< ModuleTraits > |
| Module classifying the particles of a molecular structure following their system's labels. Module classifying the particles of a molecular structure following their system's labels. More... | |
| class | T_RMSD_comb_edge_weighted_module< ModuleTraits > |
| Undefined. Undefined. More... | |
| class | T_Spatial_search_module< ModuleTraits, ApproximatedSpatialSearchEngine > |
| Module building a data base for spatial-search from an input set of points. Module building a data base for spatial-search from an input set of points. More... | |
| class | T_Union_of_balls_boundary_3_module< ModuleTraits, ExactNT > |
| Module building the boundary of the union of input 3D balls. Module building the boundary of the union of input 3D balls. More... | |
| class | T_Union_of_balls_boundary_patch_shelling_3_module< ModuleTraits, OutputArchive > |
| Module computing the shelling order forests of particles in a binding patch. Module computing the shelling order forests of particles in a binding patch. More... | |
| class | T_Union_of_balls_mesh_3_module< ModuleTraits > |
| Module computing a sampling of an input boundary of the union of 3D balls. Module computing a sampling of an input boundary of the union of 3D balls. More... | |
| class | T_Union_of_balls_surface_volume_3_module< ModuleTraits, OutputArchive > |
| Module computing the surface area and the volume of the union of an input set of 3D balls. Module computing the surface area and the volume of the union of an input set of 3D balls. More... | |
Modules framework for defining a workflow for the applications.
Modules are C++ classes instantiating C++ from the SBL core, so as to transform an input into an output. In other words, a module gives access to high level processing without having to resort to low level operations, so that an application workflow consists of interconnected modules. The Modules set of packages defines modules for selected classes from CADS, GT and CSB. In the following, three types of components are listed: