 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
 |
Structural Bioinformatics Library
Template C++ / Python API for developping structural bioinformatics applications.
|
Authors: F. Cazals and T. Dreyfus
Consider two point sets ![]() and
and ![]() of the same size; we also assume an alignment between them, meaning that the i-th point of
of the same size; we also assume an alignment between them, meaning that the i-th point of ![]() is paired with the i-th point from
is paired with the i-th point from ![]() .
.
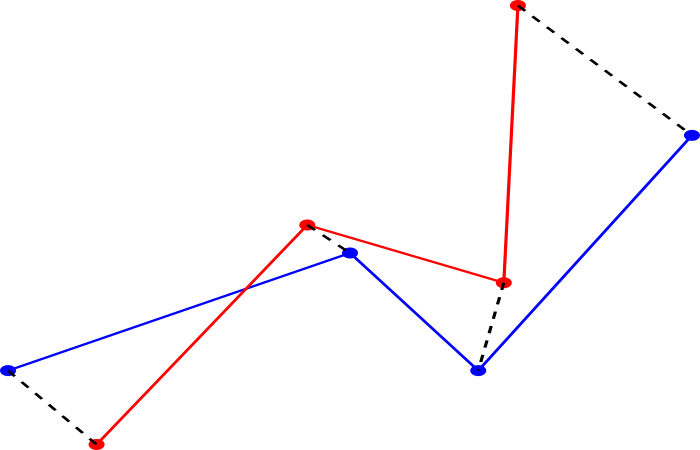
The point set registration consists on finding a spatial transformation aligning ![]() onto
onto ![]() . The registration is said rigid if the distance between all pairs of points is conserved by the transformation. This package defines a rigid registration between two set of 3D points where the spatial transformation is a composition of translations and rotations.
. The registration is said rigid if the distance between all pairs of points is conserved by the transformation. This package defines a rigid registration between two set of 3D points where the spatial transformation is a composition of translations and rotations.
Assume that we want to perform the registration of the point cloud ![]() onto the point cloud
onto the point cloud ![]() . In doing so, we shall assume that the center of mass (aka centroid) of
. In doing so, we shall assume that the center of mass (aka centroid) of ![]() is the reference center of mass, and upon applying the optimal rigid motion to
is the reference center of mass, and upon applying the optimal rigid motion to ![]() , the centroid of this modified point set will coincide with that of
, the centroid of this modified point set will coincide with that of ![]() .
.
The rigid registration is done using a linear algebra algorithm that computes the optimal rotation matrix between the two input set of points using the Singular Value Decomposition (SVD) of their covariance matrix. Basically, once centered on the origin, the covariance matrix ![]() of the two input sets of points
of the two input sets of points ![]() and
and ![]() is computed. The SVD of
is computed. The SVD of ![]() then produces orthogonal matrices representing rotations (
then produces orthogonal matrices representing rotations ( ![]() ).
).
More precisely, the algorithm is as follows :
The algorithm is coded in the class SBL::GT::T_Point_cloud_rigid_registration_3< FT >, where FT is the number type used for representing the coordinates of the points (by default, the double type).
The class can be used in two modes :
Note that the second one is suited to cases where the same transformation has to be applied to a family of point clouds.
The constructor of SBL::GT::T_Point_cloud_rigid_registration_3 takes two containers of points and computes the associated rigid transformation. Then the class provides functors and methods for the aforementioned two modes :
For both modes, the input points clouds can be represented in three different ways :
The last one is useful when the input sets of points are represented by D-dimensional points, for exemple when computing the distance between molecular conformations (see package Molecular_distances)
The following example shows how to perform a rigid registration:
The following example shows how to store the transformation defined by the rigid registration of a point cloud onto another one, so as to later use this transformation to transform any point cloud.
Note that in this example, variable points2 is registered onto points1, so that the reference centroid is that of point1. Which means that in applying the optimal rigid motion to points3, the centroid of the transformed points3 coincides with that of points1.
This package offers alors a program for performing the rigid registration between a reference set of 3D points and a collection of sets of 3D points : sbl-align-point-clouds-3.exe . An input set of 3D points is represented as a D dimensional point, where D is divisable by 3. A file listing D dimensional points is a text file where each point is represented by its dimension followed by its coordinates. Note that breaklines are not considered, so that having one D dimensional point per line, or one 3D point per line are both acceptable :
6 0 0 0 1 1 1
or
6 0 0 0 1 1 1
The program sbl-align-point-clouds-3.exe takes two files as input : the first one containing one D dimensional point representing the reference set of points, and the second containing any number of D dimensional points.